pacman::p_load(sf, tmap, sfdep, tidyverse, dplyr)In-class Exercise 5
Installing and Loading Packages
- sfdep creates a sf and tidyverse friendly interface to the package.
Data
The two data sets we will use are: - Hunan, a geospatial data in ESRI shape file format - Hunan_2012, aspatial attribute data in csv format
Import geospatial data
hunan <- st_read(dsn="data/geospatial", layer="Hunan")Reading layer `Hunan' from data source
`C:\guacodemoleh\IS415-GAA\In-class_Ex\In-class_Ex05\data\geospatial'
using driver `ESRI Shapefile'
Simple feature collection with 88 features and 7 fields
Geometry type: POLYGON
Dimension: XY
Bounding box: xmin: 108.7831 ymin: 24.6342 xmax: 114.2544 ymax: 30.12812
Geodetic CRS: WGS 84Importing aspatial data
hunan2012 <- read_csv("data/aspatial/Hunan_2012.csv")Combining aspatial fields to geospatial data
hunan_GDPPC <- left_join(hunan,hunan2012) %>%
select(1:4, 7, 15)Plotting Choropleth Map
tmap_mode("plot")
tm_shape(hunan_GDPPC) +
tm_fill(col="GDPPC",
style = "quantile",
palette = "Blues",
title = "GDPPC") +
tm_layout(main.title = "Distribution of GDP per capita by County",
main.title.position = "center",
main.title.size =1.2,
legend.height = 0.45,
legend.width = 0.35,
frame = TRUE) +
tm_borders(alpha = 0.5) +
tm_compass(type="8star", size = 2) +
tm_scale_bar() +
tm_grid(alpha = 0.2)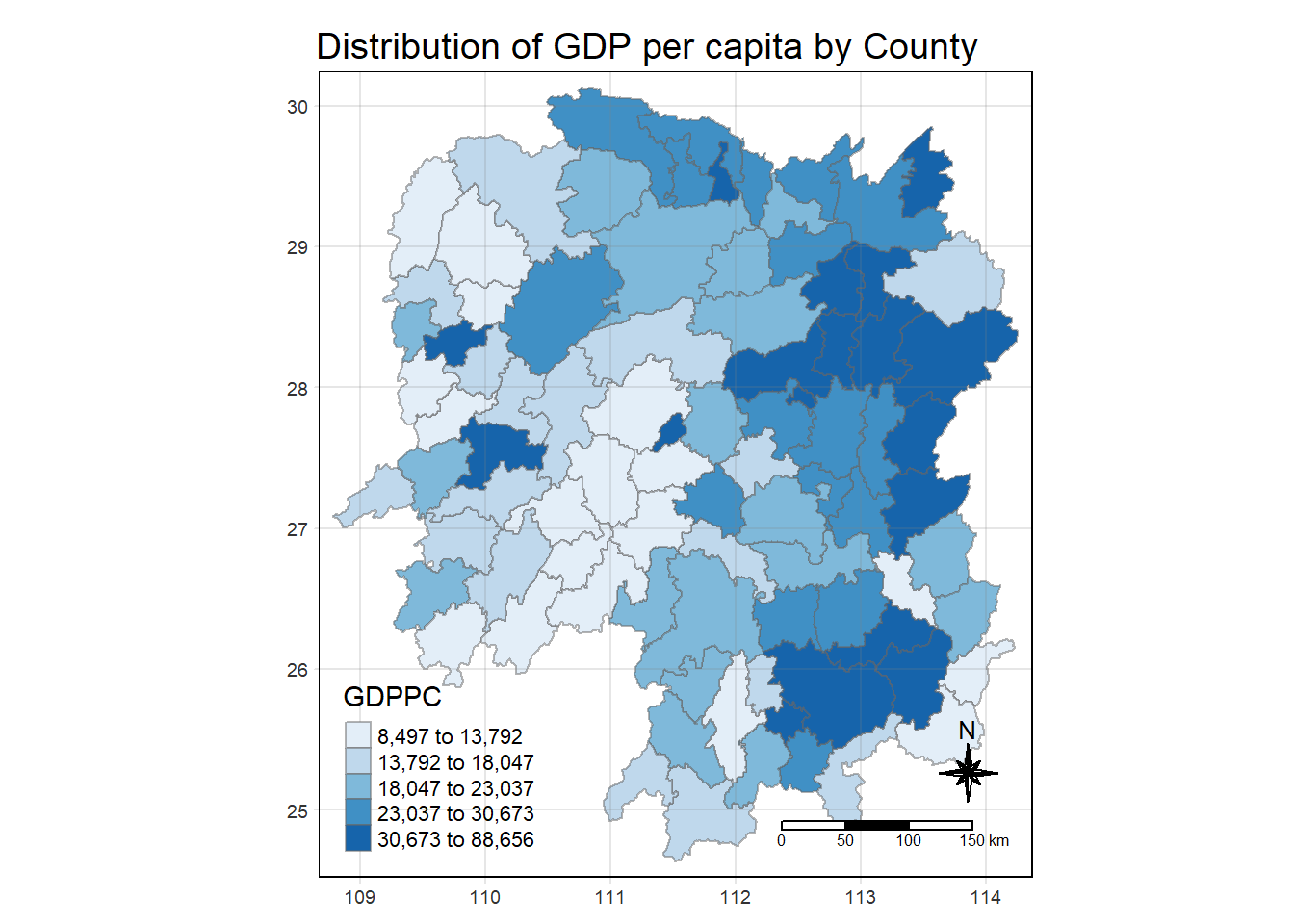
Global Measures of Spatial Association
The queen method is used to derive the contiguity weights
wm_q <- hunan_GDPPC %>%
mutate(nb = st_contiguity(geometry),
wt = st_weights(nb,
style = "W"),
.before = 1)Computing Global Moran’s I
moranI <- global_moran(wm_q$GDPPC,
wm_q$nb,
wm_q$wt)
glimpse(moranI)List of 2
$ I: num 0.301
$ K: num 7.64global_moran_test(wm_q$GDPPC,
wm_q$nb,
wm_q$wt)
Moran I test under randomisation
data: x
weights: listw
Moran I statistic standard deviate = 4.7351, p-value = 1.095e-06
alternative hypothesis: greater
sample estimates:
Moran I statistic Expectation Variance
0.300749970 -0.011494253 0.004348351 Performing Global Moran’s I permutation test
It is better to conduct permutation tests.
set.seed(1234)global_moran_perm(wm_q$GDPPC,
wm_q$nb,
wm_q$wt,
nsim = 99)
Monte-Carlo simulation of Moran I
data: x
weights: listw
number of simulations + 1: 100
statistic = 0.30075, observed rank = 100, p-value < 2.2e-16
alternative hypothesis: two.sidednsimrefers to the number of simulations- Since the p-value is < 0.05, there is sufficient evidence to reject the null hypothesis that the spatial distribution of GDP per capita resembles a random distribution, or rather the distribution is independent from spatial. Since Moran’s I statistic is > 0, we can infer that the spatial distribution shows signs of clustering.